Or: how to recontruct soft tissues in living and extinct animals!

Our co-authored paper was just published in Scientific Reports, led by Oliver E. Demuth as part of the DAWNDINOS project, led by Professor John R. Hutchinson. Find the paper here
Abstract: Biomechanical models and simulations of musculoskeletal function rely on accurate muscle parameters, such as muscle masses and lines of action, to estimate force production potential and moment arms. These parameters are often obtained through destructive techniques (i.e., dissection) in living taxa, frequently hindering the measurement of other relevant parameters from a single individual, thus making it necessary to combine multiple specimens and/or sources. Estimating these parameters in extinct taxa is even more challenging as soft tissues are rarely preserved in fossil taxa and the skeletal remains contain relatively little information about the size or exact path of a muscle. Here we describe a new protocol that facilitates the estimation of missing muscle parameters (i.e., muscle volume and path) for extant and extinct taxa. We created three-dimensional volumetric reconstructions for the hindlimb muscles of the extant Nile crocodile and extinct stem-archosaur Euparkeria, and the shoulder muscles of an extant gorilla to demonstrate the broad applicability of this methodology across living and extinct animal clades. Additionally, our method can be combined with surface geometry data digitally captured during dissection, thus facilitating downstream analyses. We evaluated the estimated muscle masses against physical measurements to test their accuracy in estimating missing parameters. Our estimated muscle masses generally compare favourably with segmented iodine-stained muscles and almost all fall within or close to the range of observed muscle masses, thus indicating that our estimates are reliable and the resulting lines of action calculated sufficiently accurately. This method has potential for diverse applications in evolutionary morphology and biomechanics
Okay… but how do you do that?
Well, first you identify the muscle’s origin and insertion. Then, follow the steps in the supplementary information using AUTODESK MAYA (free for researchers/academics/education purposes) to create a polygonal muscle, which uses a closed cylinder approach with either 8, 12 or 16 faces to describe the circumference. The cylinder’s height is arbitrarily defined depending on muscle body length. The cylinder follows the muscle’s path, as shown below for Euparkeria.
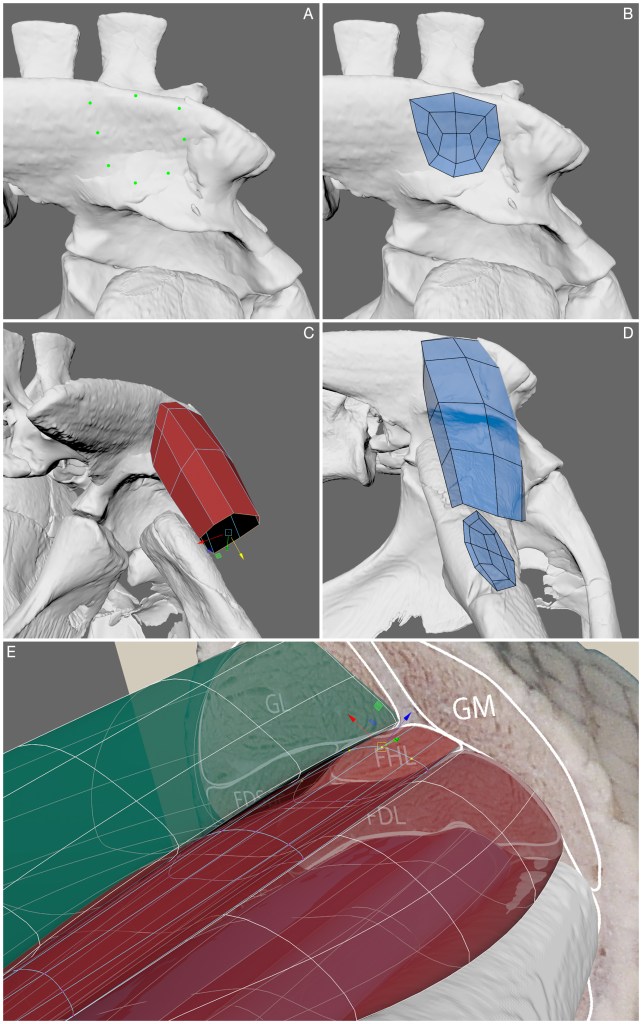
The end result is a polygon representing the volume, shape and size of a given muscle. If you wish to move beyond soft tissue reconstruction, then you can also run the provided line of action (LOA) estimation script in MAYA (uses MAYA EMBEDDED LANGUAGE) which generates a LOA, threaded through the midline of the body.
But… how do you choose the muscle path and configuration?
Well, there’s a few methods you could use! For the above, we used cross-sections of an alligator’s leg (an analagous species to Euparkeria). These cross sections helped to guide the muscle’s path and volume (see figure below) and made sure that each reconstructed muscle stayed within its boundary and reflected reality.
For living species, you can use the above method too, or instead use CT or MRI scans to guide muscle configurations, but you could also use surface scans of muscle layers! The latter is not only cheaper, but also helps to streamline the muscle modelling approach in which the user can:
(1) Collect dissection data
(2) Surface scan each topographical layer during dissection (alternatively, you could also use photogrammetry, which can be done cheaply using a smartphone with an in-built camera and free online photogrammetry software – see this wonderful blog by Dr Peter Falkingham for how-to guides and an overview of software)
(3) Use the surface scans to guide muscle paths/LOA estimation
(4) Build a subject-specific model in which dissection data AND muscle path data comes directly from the same specimen
We tried this approach on the shoulder musculature of a gorilla, as shown below.
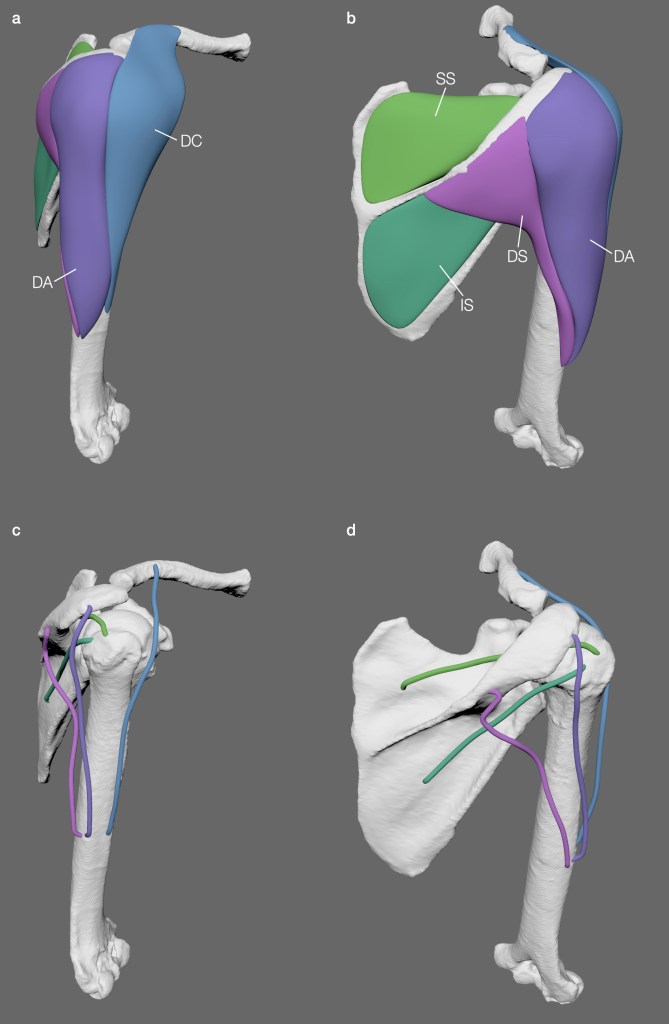


Left: shoulder musculature of a gorilla and estimated LOAs. Middle: polygonal muscle modelling approach of the gorilla surface scan data. Right: cross-section of alligator hindlimb muscles.
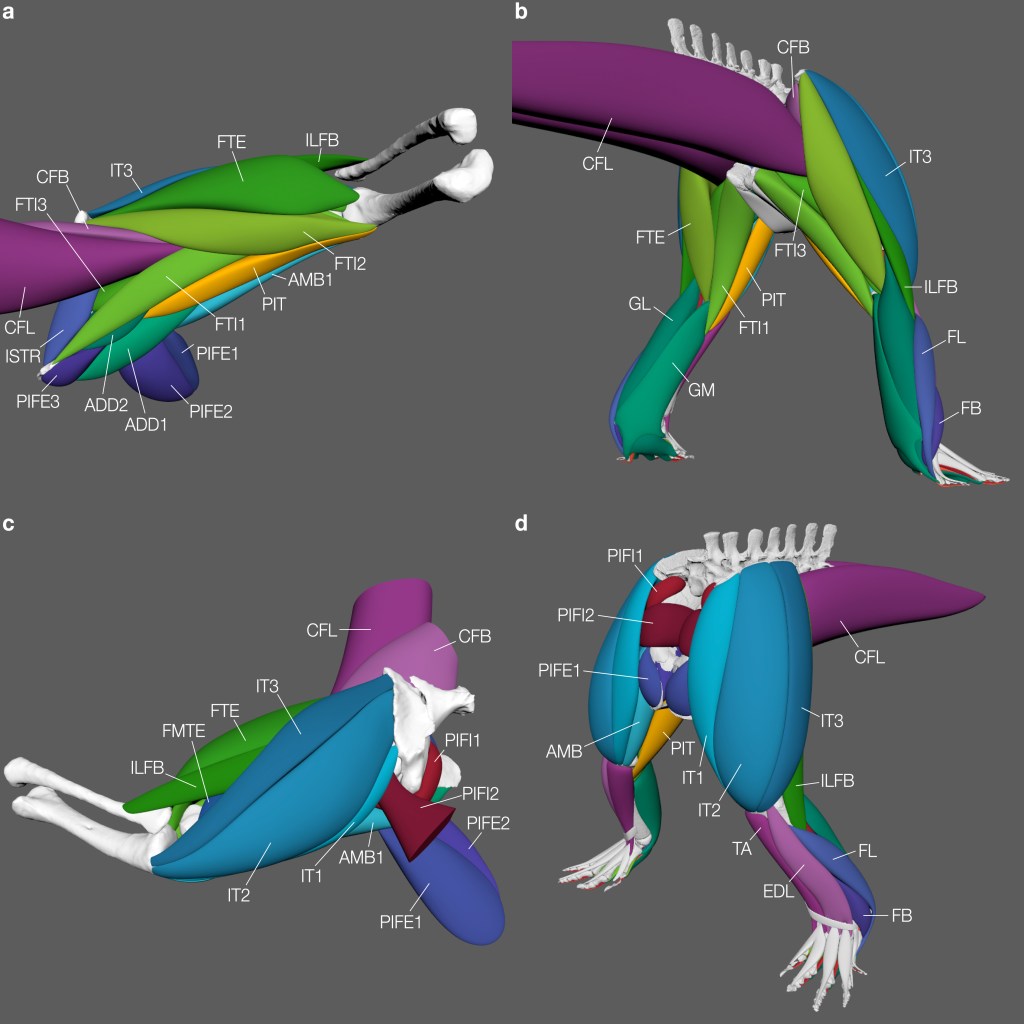
After this process, the soft tissues of the desired body segment (here, limbs) will be reconstructed! Below, we have the 3D reconstructed hindlimb and pelvic muscles of Euparkeria (see our paper for muscle abbreviations). And why would we want to do something like this? Other than it looking pretty cool, of course! For future biomechanical modelling studies… watch this space! 🙂
Leave a comment