Our new paper just landed in Integrative Organismal Biology. Go check it out here
*Copyedit version expected mid Feb 2022
This paper forms a part of my previous postdoc on the DAWNDINOS project in which we collected motion data of crocodiles using biplanar X-ray, also known as ‘XROMM’ (X-ray Reconstruction of Moving Morphology). This method involves taking a series of X-ray snapshots from at least two different angles/positions whilst an animal/object moves through a designated capture area. From this, you compile a video of the X-rays in which you can then see skeletal movement! It is pretty cool.
We placed tantalum markers inside the hindlimb of a crocodile which are quite dense and show up very well on the X-ray shadows (see the image below and try to look for black circles on the limb bones – those are the markers). These markers get tracked throughout a motion and we use those marker positions/trajectories to guide movement of the bones. For example, if the 3D position of these markers moves over there, then the respective bone on which they are attached to moves over there too.

It all sounds pretty straight forward in theory. In practise, things can go wrong during the initial setup. To place markers in the hindlimb, you first need to perform minor surgery on the animal. It’s not so simple to get straight down to the bone during surgery – there’s a lot of soft tissue in the way. And – so – mistakes happen. Sometimes beads can be misplaced, or become ‘lost’ in the body. All beads might be thought to have been successfully placed, but during the animal’s recovery period, the bead can travel elsewhere in the body! Imagine my frustration when I was tracking the XROMM data to discover that one crocodile was missing a critical bead placement in the pelvis, despite being placed in the correct position during surgery! And we only know when this happens after experimentation (which may even be after you have lost access to the animal so no further surgery and experimentation can happen). Prior to this discovery of the ‘lost’ bead, we had already established the IK rig and had tracked more than 50 trials in another animal. So this really threw a spanner in the works. Would we need to re-invent the tracking wheel? Was all data from that crocodile useless? What could we do?
And this is where our paper comes into play. What do you do when this happens? Is all lost? Is all motion data from that animal useless? Not quite!! Of course, the following will depend on the research question at hand and the intended use of the data. But if you are tracking data to be used in musculoskeletal modelling with the intention of limiting degrees of freedom, then you can use the method we have called ‘inverse kinematic (IK) marker-guided rotoscoping’*. See the below diagram to see how this method compares with other methods, such as scientific rotoscoping:
*this is because our method loses some anatomical fidelity, but may be of little to no concern for certain types of biomechanical studies
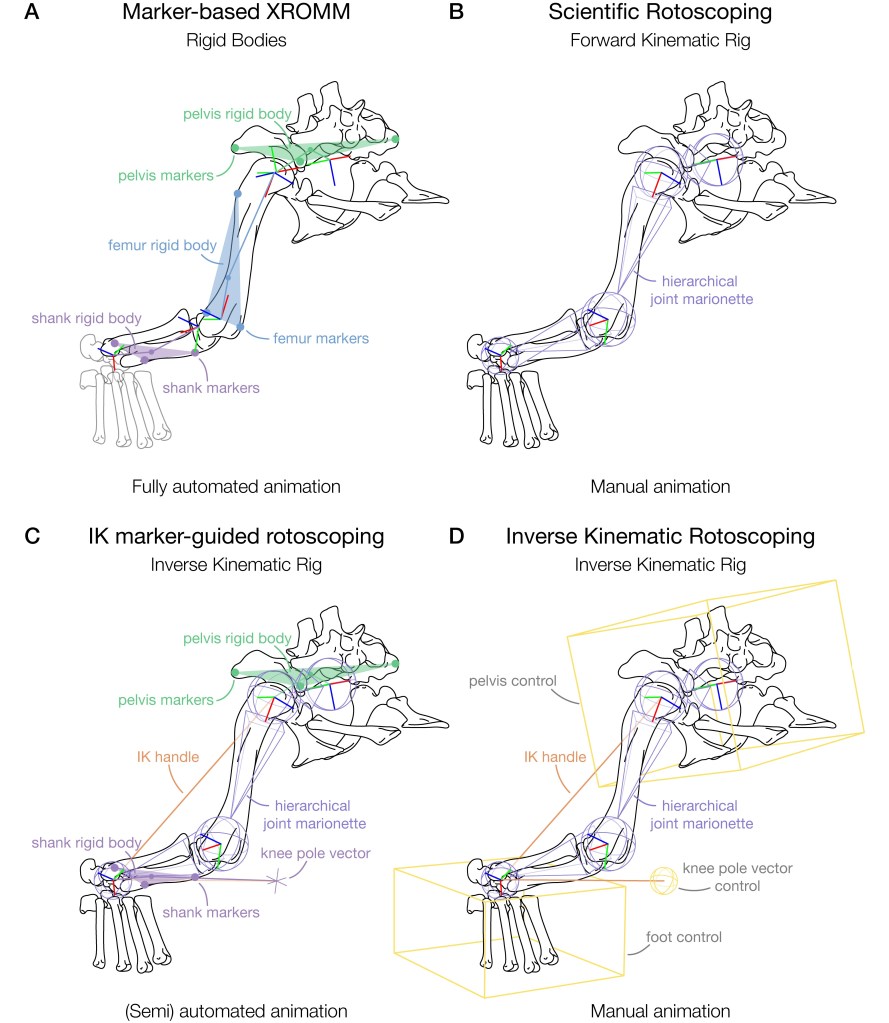
Our method combines inverse kinematic solvers with that of traditional scientific rotoscoping methods to quickly and efficiently overlay 3D bone meshes with the X-ray shadows from XROMM data. We demonstrate this method using a case study of three Nile crocodiles’ (Crocodylus niloticus) forelimbs and hindlimbs. Within these limbs, different marker configurations were used: some configurations had six markers, others had five markers and all forelimb data only had three markers*. To evaluate IK marker-guided rotoscoping, we systematically removed markers in the six-marker configuration and then tested the magnitudes of deviation in translations and rotations of the rigged setup with fewer markers versus those of the six-marker configuration. We established that IK marker-guided rotoscoping is a suitable method for ‘salvaging’ data which may have too few markers.
*The three marker configuration definitely turned out to be a bit of a headache to solve. Co-first author Oliver Demuth came up with the idea of using another IK rig controlling the first to track these trials
We illustrate how each of these setups is implemented in Autodesk Maya below:

The result of all rigs (hindlimbs and forelimbs) was a set of XROMM-informed bony motions from which the rotations of each joint and the translations and rotations of the pelvic/pectoral girdles were exported. These translations and rotations can be used to animate musculoskeletal models or conduct simulations from which we can extract biomechanical information.
Below we illustrate how accurately the different marker configurations compare to the original setup. If the tracking was perfect, then all colours would align perfectly. But we can see that there is some small deviation in the tracking which was negligible for our previous study published last year in Journal of Anatomy (check that out here):
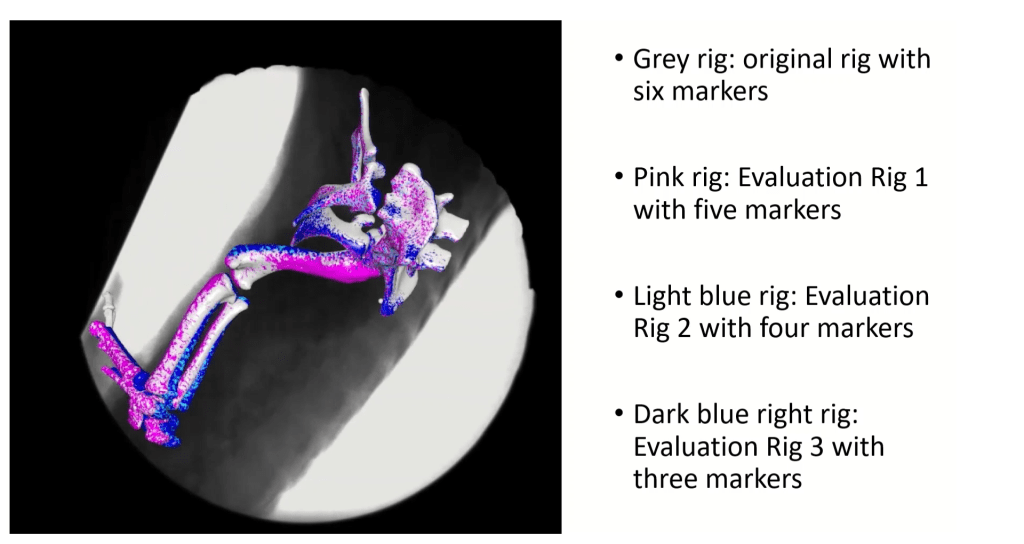
Please check out the paper to see how the method might be useful for you. Step by step instructions can be found in the supplementary material. We also provide the Maya 2019 file (.ma) which you can use as a template to set up your own IK rig in Maya.
Leave a comment